
I Let an AI Run 120 Experiments Overnight on My GPU. Here's What It Found About Biomedical NER.
A few weeks ago I stumbled onto Andrej Karpathy’s autoresearch pattern, the idea of giving an AI agent a GPU, a training script, and a goal, then letting it run experiments autonomously while you sleep. Around the same time, I noticed OpenMed had published a bunch of open-source biomedical datasets and models on HuggingFace. I wanted to play with both, so I asked Claude to help me figure out what interesting question I could explore with the medical data. It suggested looking at cross-dataset transfer: can training a model on one type of biomedical data help it recognize a completely different type?
I didn’t have a hypothesis going in. I just had tools, data, and curiosity. The answer turned out to be surprisingly asymmetric, and I think the finding is genuinely interesting.
The Setup
Named Entity Recognition (NER) is the task of finding and labeling specific things in text. In biomedical NER, you’re looking for mentions of diseases, chemicals, genes, proteins, species, etc. in scientific papers. For example:
“Treatment with aspirin reduced the risk of heart disease in patients with BRCA1 mutations.”
A good NER model would tag “aspirin” as a chemical, “heart disease” as a disease, and “BRCA1” as a gene.
The thing is, there are separate datasets for each entity type (one for chemicals, one for diseases, one for genes, etc.). They’re all biomedical text, but annotated for different things. My question was: if you first train on chemical data, does it help when you later train on disease data? And does it work the other way around?
I used five publicly available datasets:
| Dataset | What it labels |
|---|---|
| BC5CDR | Chemicals and drugs |
| NCBI Disease | Disease mentions |
| BC2GM | Genes and proteins |
| JNLPBA | DNA, RNA, proteins, cell types |
| Linnaeus | Species |
The base model is ModernBERT, a recent BERT variant. Each experiment had a fixed 5-minute training budget on an RTX 4090, and the agent decides how to split that time between pretraining on source data and fine-tuning on the target.
The Autoresearch Loop
I pointed Claude Code at the codebase with a program.md file describing the experiment protocol and went to bed. The loop is simple:
- Hypothesize: pick a curriculum (which datasets, what order, how much time on each)
- Modify the training config
- Run the experiment (5 minutes)
- Record the result
- Keep or revert: if F1 improved, commit. If not, revert and try something else.
Over two sessions, the agent ran ~120 experiments covering:
- Every source dataset individually
- Multi-stage curricula (A then B then C)
- Mixed training (A and B simultaneously)
- Time split optimization (how much time on pretraining vs fine-tuning)
- Hyperparameter sweeps (learning rate, weight decay, schedulers, batch size, etc.)
- Architecture changes (MLP heads, CRF layers)
- Multiple repeats for statistical confidence
The Main Finding: Transfer Is a One-Way Street
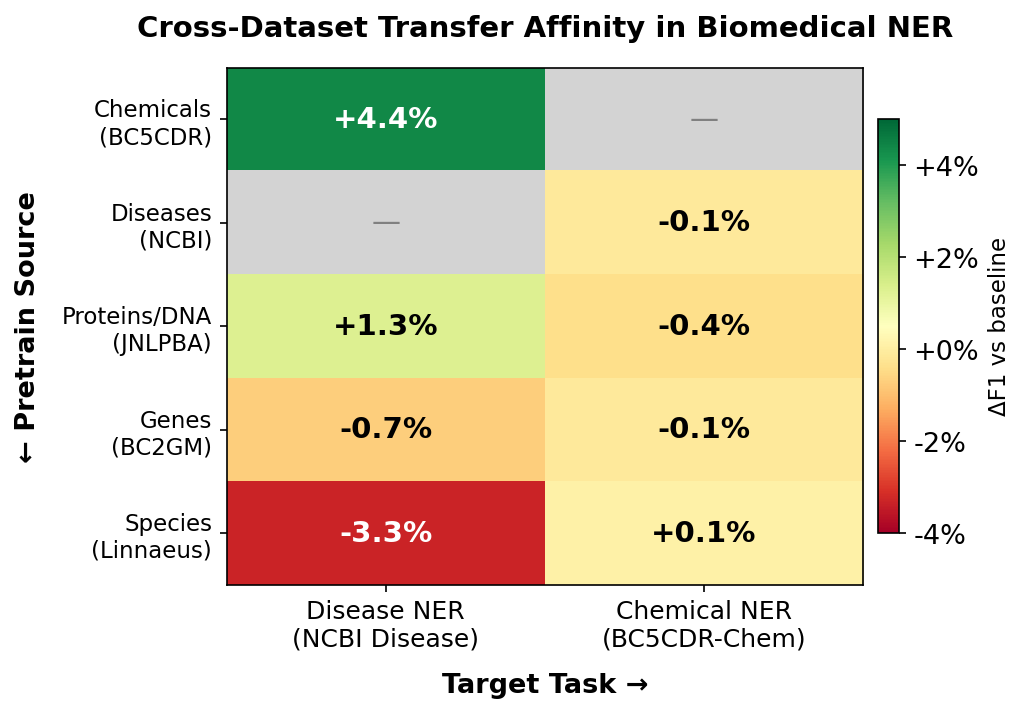
Here’s the transfer affinity matrix. Each cell shows how much pretraining on one entity type helps (or hurts) recognizing another:
| Pretrain on ↓ | Disease NER | Chemical NER |
|---|---|---|
| Chemicals (BC5CDR) | +4.4% | - |
| Diseases (NCBI) | - | -0.1% |
| Proteins/DNA (JNLPBA) | +1.3% | -0.4% |
| Genes (BC2GM) | -0.7% | -0.1% |
| Species (Linnaeus) | -3.3% | +0.1% |
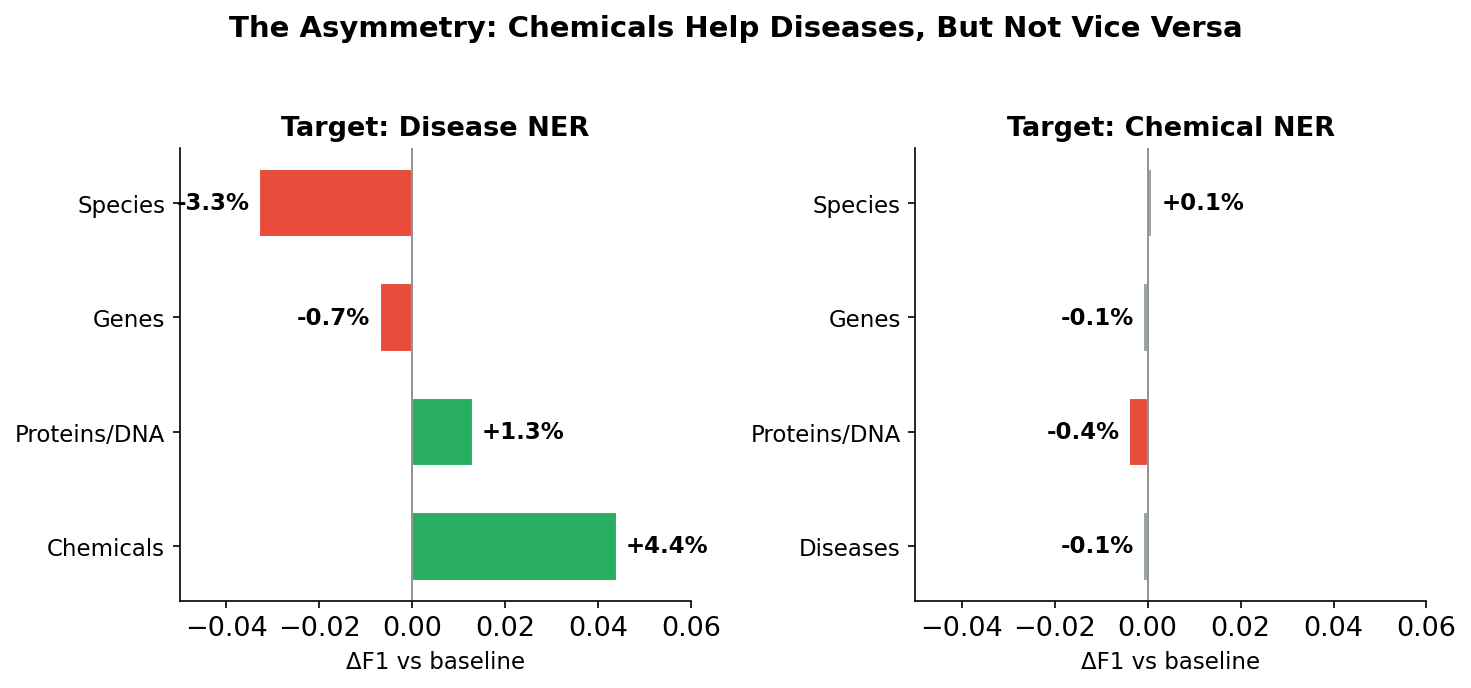
Chemical NER massively helps disease NER, but disease NER does basically nothing for chemical NER.
With curriculum optimization (multi-stage training, time split tuning), the numbers get even more striking:
| Direction | Best Improvement | How much pretrain time needed |
|---|---|---|
| Chemicals → Diseases | +5.0% | 40% of the 5-minute budget |
| Diseases → Chemicals | +1.3% | Only 15% (more hurts) |
The transfer is 3.8x stronger in one direction. I didn’t expect this at all.
Why Does This Happen?
I’m not a biologist, but here’s my engineer’s intuition for what’s going on:
1. The chemical dataset secretly teaches disease NER too. BC5CDR is annotated for both chemicals and diseases. So when you pretrain on it, the model isn’t just learning about chemicals, it’s also seeing disease mentions in context. It’s getting a head start on the exact task you’re about to fine-tune for.
2. Chemical names are more “self-contained.” Words like “aspirin”, “metformin”, or “ibuprofen” are pretty distinctive and look like chemical names regardless of context. Disease names like “diabetes”, “cancer”, or “heart failure” are trickier because they overlap with symptoms, body parts, and everyday language. Disease NER needs more contextual understanding, which is exactly what pretraining provides.
3. The datasets are different sizes. NCBI Disease has ~600 training examples. BC5CDR has ~5,000. When your target dataset is small, you benefit more from transfer learning because you have less of your own data to learn from.

Other Things I Learned
Sequential beats simultaneous. Training on chemical data then disease data works much better than mixing them together. The model benefits from focused learning on one entity type at a time.
The best curriculum has three stages. For disease NER: chemicals (25% of time) → proteins/JNLPBA (15%) → diseases (60%). The protein stage acts as a bridge between the chemical and disease domains.
Batch size matters more than you’d think in a time-limited setting. Going from batch 16 to batch 64 was a bigger improvement than most curriculum changes, simply because you process more data in the same 5 minutes. Not a deep insight, but easy to overlook.
Most hyperparameter tuning was wasted effort. I ran 50+ experiments tweaking learning rate, weight decay, warmup ratio, schedulers, dropout, gradient clipping, optimizers. The defaults (LR=5e-5, weight_decay=0.01, cosine scheduler) were basically optimal. The real gains came from curriculum design, not hyperparameter tuning.
Single-run comparisons are misleading. Early on, switching from cosine to constant_with_warmup scheduler seemed like a clear win (0.863 vs 0.861). After running each config 4-5 times, cosine actually had a higher mean (0.854 vs 0.849). The ~1% run-to-run variance from GPU non-determinism masks small differences. Always compare distributions, not single runs.
Negative transfer is real. Species NER (Linnaeus) actively hurts disease recognition (-3.3%). Not all biomedical data is created equal, and domain distance matters.
The Autoresearch Experience
Running this autonomously was both impressive and humbling. The agent systematically explored the space in a way I wouldn’t have had the patience for. 120 experiments at 5 minutes each is 10 hours of GPU time. It correctly identified dead ends quickly (gave up on species transfer after 1-2 experiments) and doubled down on promising directions (ran many variations of chemical pretraining splits).
The multi-run statistical analysis was something the agent decided to do on its own after I nudged it once. It realized that its “best result ever” (0.863) was within noise of the previous best (0.861), and from that point on it started running repeats before declaring improvements. That’s the kind of scientific rigor that’s easy to skip when you’re doing experiments manually.
What it didn’t do well: it initially assumed that any single-run improvement was real, leading to a few commits that were later reverted when variance was accounted for. And it needed a push to try truly creative ideas - most of its exploration was methodical and grid-search-style rather than hypothesis-driven.
Try It Yourself
Everything is open source: osatinsky/openmed-autoresearch
The repo includes:
- The training script and experiment protocol
- Full results from all ~120 experiments (
results.tsv) - Detailed findings (disease target, chemical target)
- Git history showing only the improvements (each commit = a new best)
To run your own exploration, you just need a GPU, the datasets, and Claude Code. Point it at program.md and let it go. You could target a different entity type, use a different base model, or extend the time budget to see if the findings hold.
What’s Next
The obvious extensions:
- Complete the matrix: run the remaining 3 target datasets (genes, proteins, species) to build a full 5x5 transfer affinity matrix
- Larger models: does the asymmetry hold with ModernBERT-large or domain-specific models like PubMedBERT?
- Longer budgets: our 5-minute constraint is artificial. With more time, do the weaker transfer directions eventually catch up?
- Cross-language: do these transfer patterns hold for biomedical NER in other languages?
If you run any of these, I’d love to hear what you find.
Code and data: huggingface.co/osatinsky/openmed-autoresearch Built with Claude Code and the autoresearch pattern by Andrej Karpathy